- Services
Services
Inoviem provides a full range of services from drug discovery to clinical development by leveraging its platforms and direct access to human pathological specimens. All the platforms and services are label free with no modification of the original molecules.
- Technology
Technology
Inoviem provides breakthrough protein technologies for each phase of the drug development process.
One of the major advantages of our label-free technologies used under physiological conditions and in human tissue – NPOT® and PIMS® – is that the test compound retains its original structure, as well as the exact same molecular structure that would be used in therapy
- Expertise
Our expertise in translational pharmacology
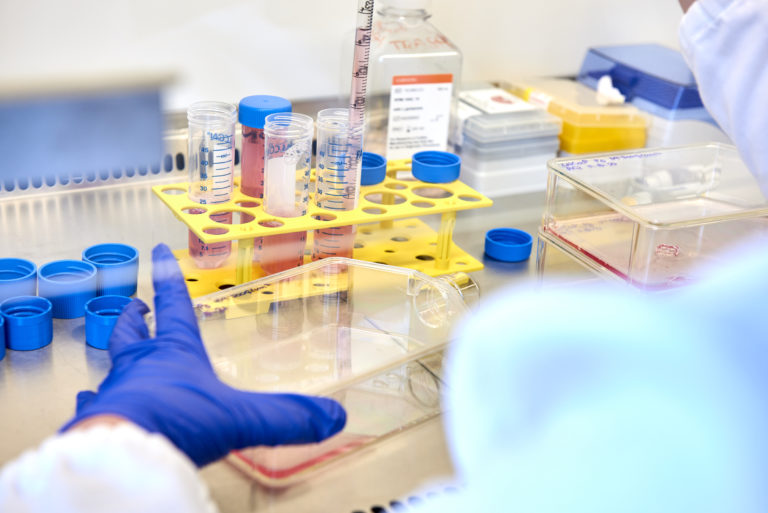
Inoviem offers translational target discovery and drug-target Mode of Action (MoA) based on proprietary label-free technologies, developed in a pathological and physiological environment, that uses human tissue.
- Knowledge centre
- About us
About us
Inoviem Scientific is a Bioanalytical R&D company established in 2011 in Strasburg – France.

Our mission
Our mission is clear: to get the right treatment to the right patients within the right therapeutic window.
- Contact us
Breakthrough protein technologies
for each phase of the drug development process
Inoviem is the only bioanalytical CRO to use label-free technologies under physiological conditions and in human tissue. NPOT® and PIMS® have proved to bring significant advantages to target identification and in confirming target engagement with compounds.
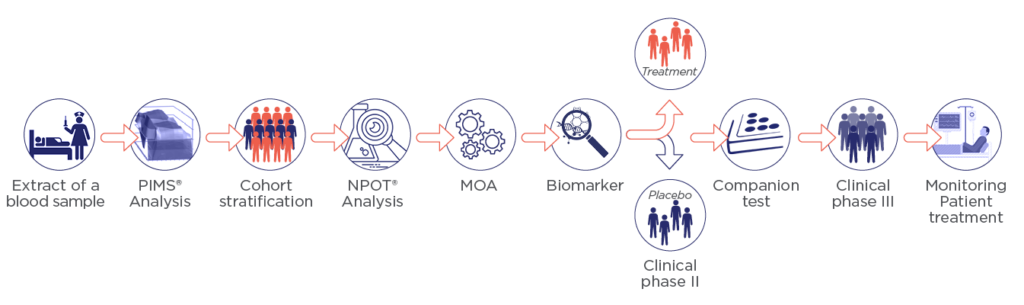
Preclinical
Identify the mode of action of your compound with NPOT® technology
NPOT technology is a unique heterogeneous assembly method for conducting extensive identification of all proteins (together with their native structure) involved in drug-target interaction.
NPOT allows us to understand the biological mechanism of a disease, a drug candidate’s mechanism of action and the origin of its side effects. This technology also enables us to identify relevant biomarkers related to a disease or associated with a drug candidate. It works by locating the target and isolating it, directly from the human tissue sample. This target is then identified and its related signaling pathway observed for pharmacological efficacy. Drug developers can then identify and validate their compound primary target and mode of action. This brings them closer to securing their drug development, funding and approval by regulatory authorities.
Conventional target deconvolution methods are unable to take into account a drug’s physiological environment, resulting in a biased ternary structure and the loss of the quaternary structure of the therapeutic target. This lost information is crucial for elucidating the biological mechanism of a disease, rational drug design, efficient structure-activity relationship studies and the investigation of aspects pertaining to target-specific toxicity and side effects. With NPOT, these key parameters are included, making your study more robust.
From preclinical to clinical
A robust solution to assure the success of your Phase II study
The individual response to a treatment can often place the success of a Phase II study in jeopardy. This occurs mainly when patient recruitment is restricted to clinical inclusion and exclusion criteria. Since we identify the responder subset within a patient cohort prior to Phase II study, the responder population is already divided into treated and placebo groups.
PIMS technology stratifies patients. NPOT identifies the mechanism of action and response at the molecular level.
Securing the development of a compound requires personalized monitoring
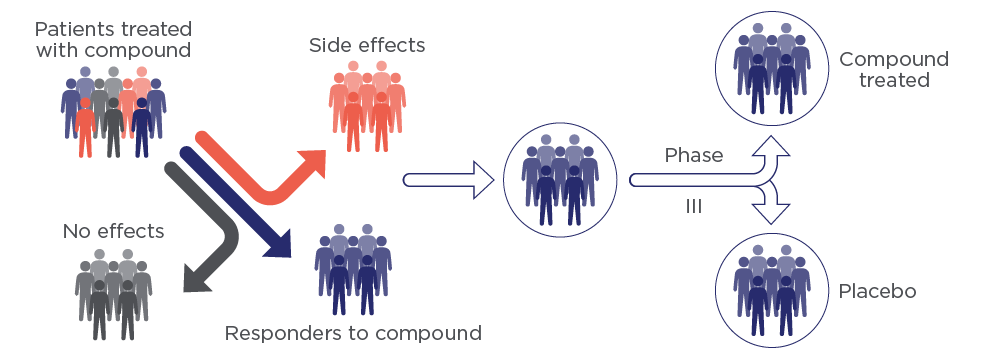
Understanding compound MoA to monitor its development
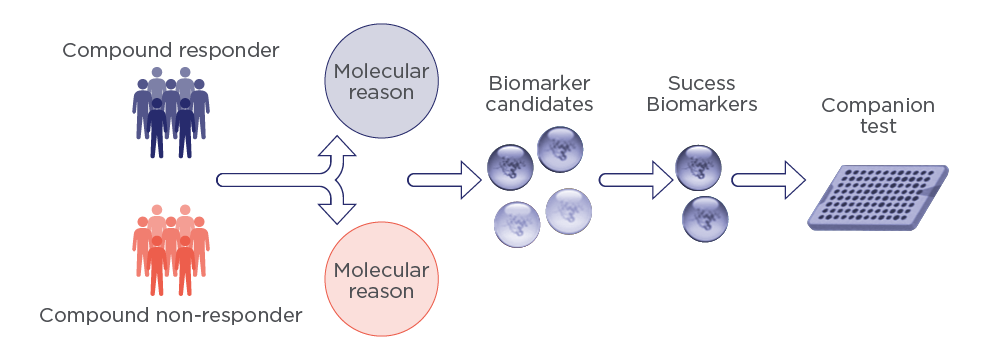
Clinical
Identify a cohort of patients responding to your therapy with PIMS® technology
PIMS is a label-free technology based on the dynamic molecular resonance of proteins and macromolecules.
PIMS is able to study protein-protein and protein-solvent interactions in multicomponent solutions. It is used to decipher responder from non-responder subjects or patients. PIMS can quickly identify the response of a tissue or cell when an exogenous molecule is administrated. It therefore reflects patients’ molecular capacity to respond to the drug’s effect and allows different subpopulations within a group to be identified in response to specific treatment.
In other words, it determines an individual fingerprint and response to a therapy.
PIMS analyses the Differential Molecular Oscillation* (DMO) and Relative Dynamic Diffraction (RDD) of a tissue in the presence of the drug candidate being tested.
This technique is used in the early stages of drug development, as well as in patient stratification.
Drug developers benefit from PIMS by being able to identify the patient group on which the compound will have a positive effect. Depending on how patients respond to different treatments on the market, PIMS can safely and accurately determine the best therapy for them. PIMS offers significant opportunities in the field of personalized medicine and biomarkers.
* Differential Molecular Oscillation (DMO) is the difference in molecular resonance as a function of time and surrounding environment in the presence of an exogenous molecule as compared to the blank. Relative Dynamic Diffraction (RDD) depends on surrounding water molecules in the presence of an exogenous molecule compared to a blank.
