- Services
Services
Inoviem provides a full range of services from drug discovery to clinical development by leveraging its platforms and direct access to human pathological specimens.
All the platforms and services are label-free with no modification of the original molecules.Unravel disease relevant Mode of Action (MoA) of your molecule directly on human tissues.
Identify and validate targets in your study model or in Human.
Thanks to our ex-vivo pharmacology expertise we identify the best disease and patients subgroups.
Using patients’ samples and PIMS® technology, we identify those individuals who can benefit best from your compound as soon as the early stage of preclinical studies.
Compounds of interest can be profiled for their drugability based on their mode of action and efficacy.
Translational pharmacology empowers your ability to identify biomarkers and advance clinical developments.
Advanced SPR and proteomics drive precise analysis of biomolecular interactions and protein landscapes.
Meticulous collection and biobanking of human samples fuel robust, translational research.
- Platforms
Platforms
Inoviem provides breakthrough protein technologies for each phase of the drug development process.
One of the major advantages of our label-free technologies used under physiological conditions and in human tissue – NPOT® and PIMS® – is that the test compound retains its original structure, as well as the exact same molecular structure that would be used in therapyPIMS®
Designed to study molecular interactions and predict therapeutic responses in biomedical research
Biomarkers discovery
Methodology for the proteomic profiling of human individuals in the context of translational evaluation of biomarkers and clinical development of new drug candidates.
- Knowledge centre
- About us
About us
Inoviem Scientific is a Bioanalytical R&D company established in 2011 in Strasburg – France.

Our mission
Our mission is clear: to get the right treatment to the right patients within the right therapeutic window.
- Contact us
PIMS®
Physiological Intermolecular Modulation Spectroscopy
PIMS® is a patented, label-free biophysical and analytical technology designed to study molecular interactions and predict therapeutic responses in biomedical research.
By leveraging near-infrared (NIR) spectroscopy, PIMS® detects subtle nanoscale modification when proteins and other biomolecules interact with drugs or other exogenous molecules. This unique approach generates dynamic “fingerprints” of a biological sample’s macromolecular structure and signalling activity, offering powerful insights into tissue response and personalized treatment outcomes.

The concept
Label-free analysis:
PIMS® operates without the need for fluorescent or radioactive labels, preserving the native state of the biological sample. It examines protein–protein and protein–solvent interactions within complex, multicomponent solutions such as blood, tissues, and PBMCs.
Water molecule resonance as a probe:
When a drug engages its target, the resulting macromolecular conformational shift alters the water resonance). This modulation provides a measurable, specific fingerprint of the treatment’s effect on molecular architecture and associated signalling pathways.
Complementary integration with NPOT®:
In tandem with the Nematic Protein Organization Technic (NPOT®), PIMS® can map the spatial organization and functional networks of proteins. This combined approach is invaluable for deciphering the mechanisms behind drug response and identifying predictive biomarkers
How PIMS® works
Dynamic fingerprinting:
PIMS® records a baseline spectral profile of a patient’s biological sample, then measures the changes after introducing a drug or compound. The resulting spectrum reflects changes in macromolecular conformation and water molecule resonance, indicating the activation (or lack thereof) of specific pharmacological signalling pathways.
Mechanism in brief:
- Sample analysis: Biological samples (e.g., blood, Plasma, Tissues, PBMCs) are challenged with a therapeutic agent.
- NIR spectroscopy: When a compound interacts with its target protein, it can induce conformational changes in the protein’s structure. These changes can alter the surrounding water molecules network, leading to a shift from high-density to low-density water structures or vice versa. PIMS measures these shifts in water molecule resonance to provide insights into the macromolecular interactions occurring within the sample.
- Signal readout: This modulation is captured as a real-time, dynamic fingerprint that differentiates responders from non-responders.
Key applications:
Personalized medicine & patient stratification:
PIMS® is used ex vivo to classify patients as good, partial, or non-responders by challenging their samples with drugs or drug combinations (e.g. lead compound plus a marketed monoclonal antibody). This stratification ensures that the most effective therapeutic strategies are selected on a patient-by-patient basis.
Disease profiling
PIMS® can distinguish between metastatic and non-metastatic patients by comparing the spectra generated with PIMS® with tumour biopsies
Biomarker development & mechanistic studies
The technology not only predicts treatment efficacy but also guides the identification of molecular networks and biomarkers. This dual readout (macromolecular conformation and water resonance) paves the way for mechanistic insights into drug action and resistance.

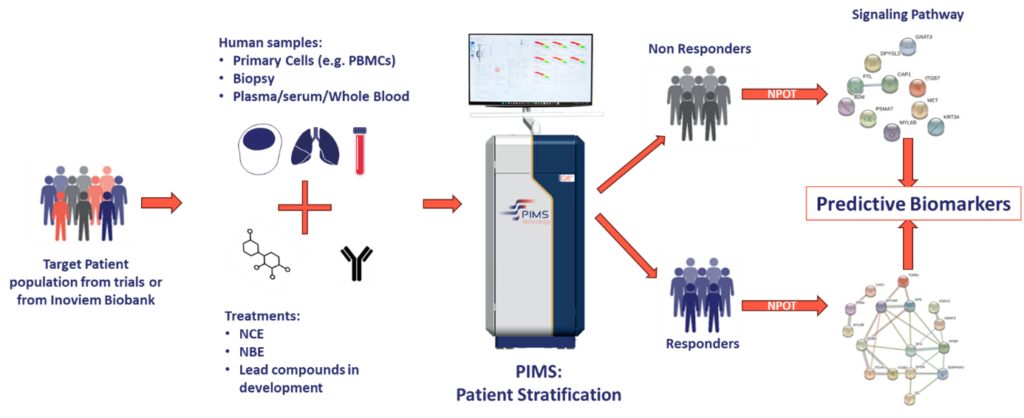
Methodology and workflow
- Sample preparation:
Biological samples are collected from patients prior to therapy. These samples—whether blood, PBMCs, or tissues—are then incubated with the drug candidate or combination of compounds. - Label-free, real-time detection:
Using NIR spectroscopy, PIMS® detects the minute shifts in water molecule resonance as the drug engages its target, triggering downstream signaling cascades. In the absence of such engagement, no significant spectral change is observed. - Data integration with NPOT®:
After stratifying patients based on their PIMS® fingerprint, NPOT® is often applied to resolve the specific signalling pathways and protein networks responsible for the observed response patterns. This two-step process provides both a predictive and mechanistic understanding of treatment efficacy.
Advantages:
Non-Invasive, holistic & rapid
The NIR approach allows for quick, non-destructive analysis without extensive sample preparation.
High specificity
PIMS® delivers precise molecular fingerprints, enabling accurate patient stratification and predictive diagnostics.
Personalized insights
By reflecting individual variations in macromolecular architecture, the technology supports personalized treatment strategies and biomarker discovery.
Why choose PIMS® at Inoviem Scientific?
At Inoviem, our commitment to cutting-edge, label-free biophysical technologies like PIMS® is at the heart of advancing personalized medicine. By providing rapid, reliable insights into how patients respond to therapeutic interventions, PIMS® is transforming drug development and clinical diagnostics—ensuring that the right treatment is delivered to the right patient.
See a PIMS® case study in Inflammatory Bowel Disease (IBD)
PIMS® has been applied to predict responses to vedolizumab therapy in patients with anti-TNF refractory IBD (including Crohn’s disease and ulcerative colitis). Clinical studies have demonstrated an 89% positive predictive value and an 82% negative predictive value, while also highlighting specific biomarkers (e.g., ITGB7, ITGAV, PF4) associated with a positive therapeutic response.
